Feature Study
Feature Review on the Use of Genomic Selection in Chicken Breeding: Current Practices and Future Prospects 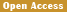
 Author
Author  Correspondence author
Correspondence author
International Journal of Molecular Zoology, 2024, Vol. 14, No. 6 doi: 10.5376/ijmz.2024.14.0032
Received: 12 Nov., 2024 Accepted: 15 Dec., 2024 Published: 26 Dec., 2024
Liang H.B., and Xuan J., 2024, Feature review on the use of genomic selection in chicken breeding: current practices and future prospects, International Journal of Molecular Zoology, 14(6): 334-343 (doi: 10.5376/ijmz.2024.14.0030)
This studyexamines the principles, methodologies, and recent advancements that underpin genomic selection (GS), a transformative approach for enhancing the efficiency of poultry breeding. Current applications of GS in improving meat and egg production traits, incorporating disease resistance, and maintaining genetic diversity are analyzed. Key enabling technologies, including high-throughput genotyping, predictive computational tools, and the integration of phenotypic and genomic data, are highlighted. A case study on the successful implementation of GS in improving disease resistance illustrates its practical impact. Challenges such as high costs, limited genomic resources, ethical concerns, and regulatory barriers are also addressed. Promising prospects are outlined for integrating GS with genome editing technologies, precision breeding, and artificial intelligence to achieve sustainable and tailored poultry production. The study emphasizes the critical role of ongoing innovation and collaboration in maximizing the potential of GS to revolutionize chicken breeding.
1 Introduction
Chicken breeding plays a pivotal role in global agriculture and food security. Poultry products, including meat and eggs, are among the most affordable and efficient sources of animal protein worldwide. The advancements in poultry breeding have significantly contributed to the increased production and efficiency of these products, thereby helping to reduce global hunger and food insecurity (Pankova and Katerinich, 2017; Athrey, 2020). The poultry industry has seen remarkable improvements in feed efficiency, production systems, and nutritional understanding, which have collectively enhanced the reliability and consistency of poultry as a food source. However, challenges such as musculoskeletal and metabolic disorders, welfare concerns, and the need to adapt to climate change threaten the long-term sustainability of this model.
Genetic improvement techniques in animal breeding have evolved significantly over the years. Traditional selective breeding methods have been the cornerstone of genetic advancements, focusing on traits such as yield, efficiency, and product quality (Chatterjee et al., 2019; Tizard et al., 2019). These methods involve mating superior individuals to enhance desirable traits in the offspring. However, the rate of genetic progress using conventional methods has been relatively slow due to the inherent nature of population and selection criteria. Recent advancements have introduced biotechnological approaches, including transgenic technology and genome editing, which offer the potential to address complex traits such as disease resistance and welfare outcomes more effectively (Doran et al., 2017). These modern techniques have shown promise in enhancing productivity and sustainability in poultry breeding (Neeteson et al., 2023).
Genomic Selection (GS) is a cutting-edge breeding technique that utilizes genomic information to predict the genetic value of individuals for specific traits. Unlike traditional methods, GS incorporates genome-wide markers to estimate breeding values, allowing for more accurate and faster selection of desirable traits. The implementation of GS in poultry breeding has led to significant improvements in traits such as growth, feed efficiency, and reproductive efficiency (Wood et al., 2019). By integrating genomic information, breeders can achieve higher precision in selection, ultimately enhancing the overall productivity and sustainability of poultry production (Neeteson et al., 2023). GS also holds potential for addressing complex traits that are difficult to improve through conventional breeding methods, such as disease resilience and welfare traits (Doran et al., 2017; Zhang and Chen, 2024).
This study attempts to explore the current practices and future prospects of genomic selection in chicken breeding, discuss the challenges and limitations associated with its implementation, and provide an overview of the historical advancements, its impact on traits such as growth, feed efficiency, and disease resistance, and potential innovations to enhance breeding efficiency and sustainability. By synthesizing the latest research and developments, it aims to offer valuable insights for researchers, breeders, and policymakers involved in poultry breeding and genetic improvement programs.
2 Fundamentals of Genomic Selection in Chicken Breeding
2.1 The concept of genomic selection: principles and methodologies
Genomic selection (GS) is a modern breeding technique that leverages high-density single nucleotide polymorphism (SNP) panels to predict the breeding values of animals based on their genomic information. This method was initially implemented in dairy cattle breeding and has since been adapted for use in poultry breeding. The core principle of GS involves the use of genome-wide markers to estimate the genetic potential of individuals, thereby enabling more accurate selection decisions at an earlier age compared to traditional methods (Wolc et al., 2016; VanRaden, 2020). The breeding value is calculated as the sum of the additive effects of alleles at all SNPs, which allows for the prediction of an individual's genetic merit for various traits (Fulton and Wolc, 2020).
2.2 Advances in genomics and bioinformatics enabling GS
Recent advancements in genomics and bioinformatics have significantly enhanced the implementation of GS in chicken breeding. High-density SNP arrays and whole-genome sequencing (WGS) have become more accessible and cost-effective, providing detailed genetic information that can be used to improve the accuracy of breeding value predictions. Techniques such as single-step genomic BLUP (ssGBLUP) have simplified the integration of genomic and pedigree data, allowing for more efficient and accurate genetic evaluations (Misztal et al., 2020). Additionally, bioinformatics tools have enabled the identification of genomic regions under selection and the annotation of candidate genes associated with economically important traits, further refining the selection process (Abdelmanova et al., 2021; Mahdabi et al., 2021).
2.3 Comparison of GS with traditional selection methods
Genomic selection offers several advantages over traditional selection methods. Traditional breeding relies heavily on phenotypic selection and pedigree information, which can be less accurate and slower due to longer generation intervals and the need for progeny testing (Wolc et al., 2016). In contrast, GS can significantly shorten generation intervals and increase the rate of genetic gain by allowing for the selection of young animals based on their genomic information (Momen et al., 2017). Moreover, GS can improve the accuracy of selection for traits with low heritability or those that are difficult to measure, such as disease resistance and reproductive traits (Gholami et al., 2015; VanRaden, 2020). However, it is important to note that while GS can enhance short-term genetic gains, it may also lead to a reduction in genetic diversity if not managed properly (Moeinizade et al., 2020).
Genomic selection has revolutionized chicken breeding by providing a more precise and efficient method for selecting superior animals. Advances in genomics and bioinformatics have played a crucial role in enabling the widespread adoption of GS, offering significant improvements over traditional selection methods. However, careful management is required to balance short-term gains with long-term genetic diversity.
3 Current Practices in Genomic Selection for Chicken Breeding
3.1 Application in meat production traits (e.g., growth rate, feed efficiency)
Genomic selection (GS) has been extensively applied to improve meat production traits in chickens, such as growth rate and feed efficiency. By utilizing high-density SNP panels, GS allows for the identification of genetic markers associated with desirable traits, leading to more accurate selection of breeding stock. For instance, studies have shown that genomic predictions using SNP panels can significantly enhance the accuracy of selecting for traits like body weight and breast muscle yield in broilers (Momen et al., 2017; Boschiero et al., 2018). Additionally, the integration of genomic data with traditional pedigree information has been shown to improve predictive correlations and reduce mean-squared predictive error, thereby optimizing breeding outcomes.
3.2 Application in egg production traits (e.g., laying rate, egg quality)
In the realm of egg production, GS has been employed to enhance traits such as laying rate and egg quality. Commercial poultry breeding companies have adopted GS to increase the rate of genetic progress for these performance traits by combining genotype and phenotype data to predict breeding values more accurately (Fulton and Wolc, 2020). For example, genomic selection has been used to identify key candidate genes associated with egg productivity and quality in different chicken breeds, thereby facilitating targeted breeding strategies (Boschiero et al., 2018; Abdelmanova et al., 2021). The use of GS in egg-type chickens has also been shown to improve the accuracy of selection and reduce the generation interval, although the scope for shortening the generation interval is less compared to meat-type chickens (Figure 1) (Wolc et al., 2016; Schmidt et al., 2023).
.png) Figure 1 Gene ontology results (Adopted from Schmidt et al., 2023) Image caption: Orange hexagons refer to GO terms and rectangles refer to proteins differentially enriched between modern (Red) and legacy (Green) chicken muscle (Adopted from Schmidt et al., 2023) |
3.3 Incorporation of disease resistance traits
Incorporating disease resistance traits into GS programs is crucial for enhancing the overall health and resilience of chicken populations. Genomic selection has been used to identify genetic markers associated with resistance to various diseases, thereby enabling the selection of individuals with superior immune responses. For instance, studies have identified candidate genes related to disease resistance, such as those involved in the immune response pathways, which can be targeted in breeding programs to improve disease resistance in chickens (Mahdabi et al., 2021; Schmidt et al., 2023). This approach not only helps in maintaining the health of the flock but also reduces the reliance on antibiotics and other treatments.
3.4 Role of GS in maintaining genetic diversity
Maintaining genetic diversity is a critical aspect of sustainable breeding programs, and GS plays a significant role in this regard. By using a wide array of genetic markers, GS allows for the selection of individuals that contribute to the genetic diversity of the population while still improving desired traits. Studies have shown that genomic selection can help in identifying and preserving genetic variants that are important for maintaining diversity, thereby preventing the loss of valuable genetic resources (Meuwissen et al., 2016; Abdelmanova et al., 2021). Additionally, the use of multi-trait prediction models that combine pedigree and genomic information can further enhance the ability to maintain genetic diversity while achieving breeding goals (Momen et al., 2017).
Current practices in genomic selection for chicken breeding have demonstrated significant advancements in improving meat and egg production traits, incorporating disease resistance, and maintaining genetic diversity. The integration of high-density SNP panels and genomic data with traditional breeding methods has led to more accurate selection and faster genetic progress. As genomic technologies continue to evolve, the future prospects for GS in chicken breeding look promising, with potential for even greater improvements in efficiency and sustainability.
4 Key Technologies Supporting Genomic Selection
4.1 High-throughput genotyping and sequencing technologies
High-throughput genotyping and sequencing technologies have revolutionized genomic selection in chicken breeding by enabling the efficient and cost-effective identification of genetic markers. The CornellGBS approach, for instance, has been optimized for chickens, allowing the successful genotyping of a large number of chickens at a cost of approximately $50 per sample. This method identified 134 528 SNPs, with 67 096 unique tags, demonstrating high performance in inferring SNPs, particularly in exonic regions and microchromosomes (Pértille et al., 2016). Additionally, high-throughput sequencing with preselection of markers has shown to be a viable alternative to SNP chips, improving the accuracy of genomic predictions in broilers (Liu et al., 2020). These technologies facilitate the large-scale application of genomic selection, enhancing the genetic improvement of economically important traits in poultry.
4.2 Computational tools and predictive models for genomic evaluations
The implementation of genomic selection in poultry relies heavily on advanced computational tools and predictive models. Single-step genomic BLUP (ssGBLUP) is a prominent method that combines genomic and pedigree relationships to create an index with all sources of information, accommodating any combination of male and female genotypes and accounting for preselection biases (Misztal et al., 2020). This method is widely used in the chicken industry due to its simplicity and accuracy. Furthermore, the development of predictive models such as BayesC and the use of high-density SNP panels have significantly improved the accuracy of genomic predictions for complex traits in broilers. These computational advancements enable more precise estimation of breeding values, thereby accelerating genetic progress in chicken breeding programs.
4.3 Integration of phenotypic and genomic data
The integration of phenotypic and genomic data is crucial for the effective application of genomic selection. By combining information from multiple genetic variants (genotypes) across the genome with trait information (phenotypes), breeding values can be predicted with greater accuracy. This approach allows for the identification of individuals with the best genetics to pass on to subsequent generations, thereby improving progeny performance (Fulton and Wolc, 2020). The use of genome-wide SNP screens to identify genomic regions under selection and key candidate genes further enhances the understanding of selection history and genomic diversity in chicken breeds, aiding in their productive breeding (Abdelmanova et al., 2021). The integration of these data types ensures a comprehensive evaluation of genetic potential, facilitating more informed selection decisions in poultry breeding programs.
High-throughput genotyping and sequencing technologies, advanced computational tools, and the integration of phenotypic and genomic data are key technologies supporting genomic selection in chicken breeding. These innovations have significantly improved the accuracy and efficiency of genetic evaluations, enabling more precise selection of breeding stock and accelerating genetic progress in the poultry industry.
5 Case Study: Successful Implementation of Genomic Selection in Poultry Breeding
5.1 Background and context of the breeding program
The implementation of genomic selection in poultry breeding has been driven by the need to enhance genetic progress and improve economically important traits. Traditional genetic improvement programs in poultry already benefit from short generation intervals, with broilers selected every six weeks and layers on an annual basis (Wolc et al., 2016). However, genomic selection offers the potential to further accelerate genetic gains by utilizing high-density SNP panels to predict breeding values more accurately and efficiently (Misztal et al., 2020).
5.2 Genomic data collection and evaluation
The methodology for implementing genomic selection in poultry involves several key steps. First, genomic data is collected using high-density SNP panels, which provide comprehensive coverage of the chicken genome (Figure 2) (Wolc et al., 2016; Li et al., 2017). For instance, in a study involving 78 chickens from 14 populations, whole-genome sequencing was performed to identify approximately 6.44 million SNPs per population. This data is then combined with phenotypic information to create genomic relationship matrices, which are used to estimate breeding values through models such as the single-step genomic BLUP (ssGBLUP) (Misztal et al., 2020). This approach integrates genomic and pedigree information, ensuring accurate and unbiased predictions of breeding values.
.png) Figure 2 Sample information and comparison of identified SNPs in each breed/population with the chicken variants database (dbSNP, Build 145) (Adopted from Li et al., 2017) Image caption: Known SNPs are SNPs already in chicken dbSNP. The map displayed here is the geographic distribution of domestic chicken populations; numbers above the dashed lines are altitudes. Red and green localities represent eight lowland and six highland chicken populations respectively, sampled in this study (Adopted from Li et al., 2017) |
5.3 Improvements in targeted traits
The implementation of genomic selection in poultry breeding has led to significant improvements in various targeted traits. For example, in a study on indigenous chicken breeding programs, the use of genomic selection resulted in a 59.3% increase in genetic gain and a 30% improvement in selection accuracy compared to conventional breeding schemes (Ndung’u et al., 2022). Additionally, genomic selection has been shown to enhance traits such as body weight, egg production, and feed efficiency in different chicken breeds (Abdelmanova et al., 2021). In broiler chickens, genetic correlations between traits like body weight, breast meat area, and egg production were effectively assessed using genomic data, leading to optimized selection strategies (Momen et al., 2017).
5.4 Lessons learned and implications for future breeding initiatives
The successful implementation of genomic selection in poultry breeding has provided several valuable lessons. One key insight is the importance of integrating both genomic and pedigree information to achieve the most accurate predictions of breeding values (Momen et al., 2017; Misztal et al., 2020). Additionally, the use of high-density SNP panels and comprehensive genomic data collection is crucial for identifying key candidate genes and genomic regions under selection (Qanbari et al., 2015; Abdelmanova et al., 2021). Future breeding initiatives can benefit from these findings by continuing to leverage genomic selection to enhance genetic progress, improve economically important traits, and maintain genetic diversity in poultry populations (Mahdabi et al. 2021).
The case study of genomic selection in poultry breeding demonstrates its effectiveness in improving targeted traits and accelerating genetic progress. By integrating genomic and phenotypic data, breeders can make more informed selection decisions, ultimately leading to more productive and efficient poultry populations. The lessons learned from these implementations highlight the potential for genomic selection to revolutionize poultry breeding and set the stage for future advancements in the field.
6 Challenges and Limitations of Genomic Selection in Chicken Breeding
6.1 High costs of genotyping and computational infrastructure
The implementation of genomic selection in chicken breeding is often hindered by the high costs associated with genotyping and the necessary computational infrastructure. High-density SNP panels, while effective, are expensive, limiting their widespread use in breeding programs. For instance, the cost of using a 600K Affymetrix Axiom high-density SNP chip is prohibitively high for genotyping all selection candidates, necessitating the development of lower-cost alternatives such as low-density SNP chips and imputation methods (Herry et al., 2020). Additionally, the computational resources required to process and analyze large genomic datasets further add to the overall costs, making it challenging for smaller breeding operations to adopt these technologies (Pértille et al., 2016).
6.2 Limited genomic resources in non-model chicken breeds
Another significant challenge is the limited availability of genomic resources for non-model chicken breeds. Most genomic selection efforts have focused on commercial breeds, leaving indigenous and less common breeds with fewer genomic tools and resources. This disparity can hinder the genetic improvement of these breeds, which may possess valuable traits for specific environments or production systems. For example, studies have shown that while commercial breeds have been extensively genotyped and analyzed, indigenous breeds often lack comprehensive genomic data, making it difficult to apply genomic selection effectively (Ndung’u et al., 2022). This limitation underscores the need for more inclusive genomic research that encompasses a broader range of chicken breeds.
6.3 Ethical and welfare concerns
The application of genomic selection in chicken breeding also raises ethical and welfare concerns. Intensive selection for specific traits can lead to unintended consequences, such as increased susceptibility to diseases or poor welfare outcomes. For instance, the selection for rapid growth in broilers has been associated with health issues like skeletal deformities and cardiovascular problems. Moreover, the focus on production traits can sometimes overshadow the importance of maintaining genetic diversity and overall animal well-being. Ethical considerations must be integrated into breeding programs to ensure that the welfare of the animals is not compromised in the pursuit of genetic gains (Marchesi et al., 2017).
6.4 Regulatory and industry-level adoption challenges
Finally, the adoption of genomic selection at the regulatory and industry levels presents its own set of challenges. Regulatory frameworks for the use of genomic technologies in animal breeding are still evolving, and there can be significant variability between regions. This lack of standardized regulations can create barriers to the implementation of genomic selection practices. Additionally, industry stakeholders may be hesitant to adopt new technologies due to the perceived risks and uncertainties associated with them. For example, the poultry industry has traditionally relied on well-established breeding practices, and transitioning to genomic selection requires a shift in both mindset and operational procedures (Wolc et al., 2016; Misztal et al., 2020). Overcoming these challenges will require concerted efforts to educate stakeholders and develop clear regulatory guidelines that support the integration of genomic technologies into breeding programs.
The use of genomic selection in chicken breeding faces several challenges, including high costs of genotyping and computational infrastructure, limited genomic resources for non-model breeds, ethical and welfare concerns, and regulatory and industry-level adoption hurdles. Addressing these challenges will be crucial for the successful implementation and widespread adoption of genomic selection in the poultry industry (Teng et al., 2019).
7 Future Prospects of Genomic Selection in Chicken Breeding
7.1 Integration of GS with genome editing technologies (e.g., CRISPR/Cas9)
The integration of genomic selection (GS) with genome editing technologies such as CRISPR/Cas9 holds significant promise for the future of chicken breeding. CRISPR/Cas9 allows for precise modifications at specific genomic loci, which can be used to introduce desirable traits identified through GS. For instance, the CRISPR/Cas9 system has been successfully used to create highly productive chickens with improved traits by targeting specific genes (Larkina et al., 2021). This combination can accelerate the breeding process by directly editing the genes associated with favorable traits, thereby enhancing the efficiency and effectiveness of GS.
7.2 Prospects for precision breeding and tailored poultry lines
Precision breeding aims to develop poultry lines that are tailored to specific production goals or environmental conditions. By leveraging GS, breeders can predict the genetic potential of chickens with high accuracy, allowing for the selection of individuals that best meet the desired criteria. This approach can optimize traits such as growth rate, feed efficiency, and disease resistance (Wolc et al., 2016; Ndung’u et al., 2022). The ability to tailor poultry lines to specific needs not only improves productivity but also enhances animal welfare and sustainability in poultry production.
7.3 Leveraging big data and artificial intelligence in genomic evaluations
The use of big data and artificial intelligence (AI) in genomic evaluations is set to revolutionize chicken breeding. Advanced computational tools and machine learning algorithms can handle the vast amounts of data generated by high-density SNP panels and whole-genome sequencing (Montesinos-López et al., 2023). These technologies can improve the predictive accuracy of genomic estimated breeding values (GEBVs) by identifying complex patterns and interactions within the genomic data. AI-driven models can also facilitate the integration of environmental and phenotypic data, further enhancing the precision of GS (Wang et al., 2018; Li et al., 2023).
7.4 Global collaboration and resource sharing for GS research
Global collaboration and resource sharing are crucial for advancing GS research in chicken breeding. By pooling genetic and phenotypic data from diverse populations, researchers can create more robust reference populations, which are essential for accurate genomic predictions (Meuwissen et al., 2016; Tan et al., 2017). International partnerships can also facilitate the exchange of technological advancements and best practices, accelerating the implementation of GS across different regions and breeding programs. Collaborative efforts can lead to the development of standardized protocols and shared databases, ultimately benefiting the global poultry industry.
The future of genomic selection in chicken breeding is promising, with significant advancements expected through the integration of genome editing technologies, precision breeding, big data, and AI. Global collaboration will play a pivotal role in maximizing the potential of GS, leading to more efficient and sustainable poultry production.
8 Concluding Remarks
Genomic selection (GS) has revolutionized chicken breeding by leveraging high-density SNP panels to enhance the accuracy of breeding value predictions and reduce generation intervals. Traditional genetic improvement programs in poultry already benefit from short generation intervals, but GS has further optimized these processes, particularly in layers where some scope for shortening the generation interval exists. The implementation of GS has been facilitated by technological advancements such as single-step genomic BLUP (ssGBLUP), which integrates genomic and pedigree data to provide accurate breeding values. Additionally, the identification of genomic regions under selection in various chicken breeds has provided valuable insights into the genetic basis of economically important traits, aiding in the development of more efficient breeding strategies.
Continued innovation and research in GS are crucial for maintaining and enhancing the genetic progress in chicken breeding. The development of new genomic selection methods, such as the complementarity-based selection strategy (CBS), aims to balance short-term genetic gains with long-term genetic diversity and growth potential. Moreover, the integration of whole-genome sequence data and the exploration of multiple-trait genomic selection can further improve the accuracy and efficiency of breeding programs. Addressing challenges such as the reduction in genetic variances due to the Bulmer effect and the need for new validation procedures unaffected by selection will be essential for the sustained success of GS.
Genomic selection holds transformative potential for sustainable poultry production by enabling more precise and efficient breeding practices. The ability to predict breeding values with high accuracy early in life allows for the selection of superior individuals, thereby accelerating genetic progress and improving productivity. The identification of key candidate genes and genomic regions under selection provides a deeper understanding of the genetic architecture of important traits, facilitating targeted breeding efforts. As the poultry industry continues to adopt and refine GS technologies, the potential for sustainable and productive poultry production will be significantly enhanced, ensuring the long-term viability and success of the industry.
Acknowledgments
We are grateful to Dr. Yang for critically reading the manuscript and providing valuable feedback that improved the clarity of the text. We express our heartfelt gratitude to the two anonymous reviewers for their valuable comments on the manuscript.
Conflict of Interest Disclosure
The authors affirm that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Abdelmanova A., Dotsev A., Romanov M., Stanishevskaya O., Gladyr E., Rodionov A., Vetokh A., Volkova N., Fedorova E., Gusev I., Griffin D., Brem G., and Zinovieva N., 2021, Unveiling comparative genomic trajectories of selection and key candidate genes in egg-type russian white and meat-type white cornish chickens, Biology, 10(9): 876.
https://doi.org/10.3390/biology10090876
Athrey G., 2020, Poultry genetics and breeding, Animal Agriculture, 2020: 317-330.
https://doi.org/10.1016/b978-0-12-817052-6.00018-5
Boschiero C., Moreira G., Gheyas A., Godoy T., Gasparin G., Mariani P., Paduan M., Cesar A., Ledur M., and Coutinho L., 2018, Genome-wide characterization of genetic variants and putative regions under selection in meat and egg-type chicken lines, BMC Genomics, 19: 1-18.
https://doi.org/10.1186/s12864-018-4444-0
Chatterjee R., Bhattacharya T., and Paul S., 2019, Breeding poultry for improved input use efficiency and nutrient quality of products, Indian Journal of Genetics and Plant Breeding, 79(Sup-01): 204-207.
https://doi.org/10.31742/ijgpb.79s.1.10
Doran T., Challagulla A., Cooper C., Tizard M., and Jenkins K., 2017, Genome editing in poultry-opportunities and impacts, National Institutes of Bioscience Journal, 1: 1-15.
https://doi.org/10.2218/NATLINSTBIOSCI.1.2016.1742
Fulton J., and Wolc A., 2020, Application of genomic selection in commercial egg-type populations, Advances in Poultry Genetics and Genomics, 2020: 403-420.
https://doi.org/10.19103/as.2020.0065.21
Gholami M., Reimer C., Erbe M., Preisinger R., Weigend A., Weigend S., Servin B., and Simianer H., 2015, Genome scan for selection in structured layer chicken populations exploiting linkage disequilibrium information, PLoS One, 10(7): e0130497.
https://doi.org/10.1371/journal.pone.0130497
Herry F., Druet D., Hérault F., Varenne A., Burlot T., Roy L., and Allais S., 2020, Interest of using imputation for genomic evaluation in layer chicken, Poultry Science, 99: 2324-2336.
https://doi.org/10.1016/j.psj.2020.01.004
Larkina T., Krutikova A., Peglivanyan G., Shcherbakov Y., and Barkova O., 2021, Development of optimal technological approaches for obtaining PGCs in Pushkin breed chickens for further transformation by the CRISPR / Cas9 system, The FASEB Journal, 35(S1): 20-76.
https://doi.org/10.1096/FASEBJ.2021.35.S1.05031
Li D., Che T., Chen B., Tian S., Zhou X., Zhang G., Li M., Gaur U., Li Y., Luo M., Zhang L., Xu Z., Zhao X., Yin H., Wang Y., Jin L., Tang Q., Xu H., Yang M., Zhou R., Li R., Zhu Q., and Li M., 2017, Genomic data for 78 chickens from 14 populations, GigaScience, 6: 1-5.
https://doi.org/10.1093/gigascience/gix026
Li T., Jiang S., Fu R., Wang X., Cheng Q., and Jiang S., 2023, IP4GS: Bringing genomic selection analysis to breeders, Frontiers in Plant Science, 14: 1131493.
https://doi.org/10.3389/fpls.2023.1131493
Liu T., Luo C.J., Wang Y., Shu D., Su G., and Qu H., 2020, High-throughput sequencing with the preselection of markers is a good alternative to SNP chips for genomic prediction in broilers, Frontiers in Genetics, 11: 108.
https://doi.org/10.3389/fgene.2020.00108
Mahdabi E., Esmailizadeh A., Mehrgardi A., and Fozi A., 2021, A genome-wide scan to identify signatures of selection in two Iranian indigenous chicken ecotypes, Genetics, Selection, Evolution, 53: 1-16.
https://doi.org/10.1186/s12711-021-00664-9
Marchesi J., Buzanskas M., Cantao M., Ibelli A., Peixoto J., Joaquim L., Moreira G., Godoy T., Sbardella A., De Figueiredo E., Coutinho L., Munari D., and Ledur M., 2017, Relationship of runs of homozygosity with adaptive and production traits in a paternal broiler line, Animal, 12(6): 1126-1134.
https://doi.org/10.1017/S1751731117002671
Meuwissen T., Hayes B., and Goddard M., 2016, Genomic selection: a paradigm shift in animal breeding, Animal Frontiers, 6: 6-14.
https://doi.org/10.2527/AF.2016-0002
Misztal I., Lourenco D., and Legarra A., 2020, Current status of genomic evaluation, Journal of Animal Science, 98(4): skaa101.
https://doi.org/10.1093/jas/skaa101
Moeinizade S., Wellner M., Hu G., and Wang L., 2020, Complementarity‐based selection strategy for genomic selection, Crop Science, 60: 149-156.
https://doi.org/10.1002/csc2.20070
Momen M., Mehrgardi A., Sheikhy A., Esmailizadeh A., Fozi M., Kranis A., Valente B., Rosa G., and Gianola D., 2017, A predictive assessment of genetic correlations between traits in chickens using markers, Genetics, Selection, Evolution, 49: 1-14.
https://doi.org/10.1186/s12711-017-0290-9
Montesinos-López O., Crespo-Herrera L., Pierre C., Bentley A., De La Rosa-Santamaria R., Ascencio-Laguna J., Agbona A., Gerard G., Montesinos‐López A., and Crossa J., 2023, Do feature selection methods for selecting environmental covariables enhance genomic prediction accuracy?, Frontiers in Genetics, 14: 1209275.
https://doi.org/10.3389/fgene.2023.1209275
Ndung’u C., Muasya T., and Okeno T., 2022, Optimization of response to selection using genomic selection in indigenous chicken breeding programmes, South African Journal of Animal Science, 51(6): 723-734.
https://doi.org/10.4314/sajas.v51i6.5
Neeteson A., Avendaño S., Koerhuis A., Duggan B., Souza E., Mason J., Ralph J., Rohlf P., Burnside T., Kranis A., and Bailey R., 2023, Evolutions in commercial meat poultry breeding, Animals, 13(19): 3150.
https://doi.org/10.3390/ani13193150
Pankova S., and Katerinich O., 2017, Efficiency of using the new domestic meat-egg hybrid for the production of food eggs in household farms, 4: 47-51.
https://doi.org/10.15407/agrisp4.02.047
Pértille F., Guerrero-Bosagna C., Silva V., Boschiero C., De Ribamar Da Silva Nunes J., Ledur M., Jensen P., and Coutinho L., 2016, High-throughput and cost-effective chicken genotyping using next-generation sequencing, Scientific Reports, 6(1): 26929.
https://doi.org/10.1038/srep26929
Qanbari S., Seidel M., Strom T., Mayer K., Preisinger R., and Simianer H., 2015, Parallel selection revealed by population sequencing in chicken, Genome Biology and Evolution, 7: 3299-3306.
https://doi.org/10.1093/gbe/evv222
Schmidt C., Kim D., Pendarvis G., Abasht B., and McCarthy F., 2023, Proteomic insight into human directed selection of the domesticated chicken Gallus gallus, PLoS One, 18(8): e0289648.
https://doi.org/10.1371/journal.pone.0289648
Tan C., Bian C., Yang D., Li N., Wu Z., and Hu X., 2017, Application of genomic selection in farm animal breeding, Yi chuan = Hereditas, 39(11): 1033-1045.
https://doi.org/10.16288/j.yczz.17-286
Teng J., Gao N., Zhang H., Li X., Li J., Zhang H., Zhang X., and Zhang Z., 2019, Performance of whole genome prediction for growth traits in a crossbred chicken population, Poultry Science, 98: 1968-1975.
https://doi.org/10.3382/ps/pey604
Tizard M., Jenkins K., Cooper C., Woodcock M., Challagulla A., and Doran T., 2019, Potential benefits of gene editing for the future of poultry farming, Transgenic Research, 28: 87-92.
https://doi.org/10.1007/s11248-019-00139-0
VanRaden P., 2020, Symposium review: how to implement genomic selection, Journal of Dairy Science, 103(6): 5291-5301.
https://doi.org/10.3168/jds.2019-17684
Wang X., Wang X., Xu Y., Hu Z., and Xu C., 2018, Genomic selection methods for crop improvement: current status and prospects, The Crop Journal, 6(4): 330-340.
https://doi.org/10.1016/J.CJ.2018.03.001
Wolc A., Kranis A., Arango J., Settar P., Fulton J., O'sullivan N., Avendano A., Watson K., Hickey J., Campos G., Fernando R., Garrick D., and Dekkers J., 2016, Implementation of genomic selection in the poultry industry, Animal Frontiers, 6: 23-31.
https://doi.org/10.2527/AF.2016-0004
Wood B., Emamgholi-Begli H., Abdella E., Vanderhout R., and Baes C., 2019, 181 enhancing the efficiency of poultry production by optimising selection objectives and breeding strategies, Journal of Animal Science, 97: 182-183. https://doi.org/10.1093/jas/skz258.376.
Zhang S.P., and Chen H.Y., 2024, Research on the threat of H5N1 avian influenza virus to chicken health and its molecular mechanisms, International Journal of Molecular Veterinary Research, 14(1): 23-31.
https://doi.org/10.5376/ijmvr.2024.14.0004

. PDF(456KB)
. FPDF(win)
. FPDF(mac)
. HTML
. Online fPDF
Associated material
. Readers' comments
Other articles by authors
. Hongbo Liang
. Jia Xuan
Related articles
. Chicken breeding
. Genomic selection
. Genetic improvement
. Poultry production
. Sustainable agriculture
Tools
. Email to a friend
. Post a comment


